How one lab's decadeslong search for the 'holy grail' of sickle cell led to the first CRISPR therapy
In a 2008 issue of Science, just a page apart, were the two crucial discoveries that would form the cure for sickle cell disease.
Stuart Orkin, a hematologist at Boston Children’s Hospital, had published work identifying the genetic switch that could turn on backup copies of life-sustaining hemoglobin proteins in sickle cell patients. And in the very next article, a pair of microbiologists from Illinois described a system called CRISPR — the tool that could flip that switch, but a mere curiosity at the time.
“Worlds collided, right on paper,” Orkin recalled in his seventh floor office of the hospital last week.
The research was hailed as the “holy grail” for sickle cell, a disease that predominantly afflicts Black people and for which new treatments have been slow to come despite understanding its root cause for nearly 75 years.
“We all knew that fetal hemoglobin was beneficial if you had more of it in sickle cell or in thalassemia,” Orkin said. But for decades, scientists couldn’t figure out what turned production of fetal hemoglobin on and off. “We didn’t know what we were looking for.”
It would take years for those two discoveries to converge, culminating a decade and a half later in the first-ever commercial gene editing therapy, already approved in the UK and was just approved in the US today.
And while the drug itself belongs to CRISPR Therapeutics and Vertex Pharmaceuticals — which made, tested, and commercialized it — Casgevy is in many ways Orkin’s brainchild, and the outcome of years of work by teams around the globe who pieced together the targets and tools that would make it possible.
A cause without a cure
Sickle cell disease is characterized by red blood cells that are malformed into a sickle shape, limiting their ability to transport oxygen. But they are not that way at birth: In 1948, Brooklyn pediatrician Janet Watson noticed that babies who would later go on to develop sickle cell were born with mostly healthy blood cells.
A year later, in 1949, chemist Linus Pauling discovered that the structure of hemoglobin proteins were different in people with sickle cell. The study was celebrated as the first molecular description of a disease, and led biologist Vernon Ingram to pinpoint the precise mutations that cause sickling in 1956.
Watson hypothesized that fetal hemoglobin was resistant to sickling. And there were some people who carried genes for sickle cell but whose blood cells, for reasons unknown, never stopped making fetal hemoglobin.
The discoveries captivated generations of hematologists, but would frustrate them as well. Sickle cell became a textbook example of a genetic disease, where a single typo creates enormous trouble. Even when CRISPR gene editing debuted a decade ago, the tool was better at silencing unwanted genes than fixing broken ones. And cutting out the typo would only mangle hemoglobin further.
For decades, the solution felt like it was in plain sight. Turn back on fetal hemoglobin, and you could give patients a normal life. And there was pharmaceutical progress on that front: Hydroxyurea, a drug used to slow the growth of cancer, in the early 1980s was shown to trigger modest fetal hemoglobin production in monkeys, and the drug was eventually approved for treating sickle cell patients in 1998. But it doesn’t completely address the disease, doesn’t work for everyone, and can cause side effects like nausea and vomiting.
A rush to clone and sequence genes in the 1980s fueled a search for the enigmatic switch. Scientists stared at the genetic code of different forms of hemoglobin looking for clues. But it was unknown if a single protein controlled the process, or if there was a more complex circuit involved.
“During the doldrums, we thought there were probably five to 10 factors, and what was the hope we’re ever going to figure this out?” Orkin said. “It was a frustrating era.”
Stuck, he moved onto other problems and the search for the switch slowly fizzled out.
The search for the switch
Almost 20 years later in 2004, an MD-PhD student named Vijay Sankaran was interested in sickle cell disease and had seen hydroxyurea help, but also knew its limitations. Orkin’s lab had long since given up on finding the fetal hemoglobin switch, and Sankaran wanted to resume the search.
Orkin offered him a dose of reality. “That’s great,” Orkin told him. “It’s really the most important problem we can work on. But most of the people who worked on it have either failed, they’ve gone on to other projects, or they’re dead.”
But one thing had changed. The recent completion of the Human Genome Project was propelling genetic sleuthing into a new era. One of Sankaran’s professors, the future Vertex executive David Altshuler, preached that faster and cheaper DNA sequencing would allow scientists to compare hundreds, maybe thousands, of genomes to fish out genetic variants linked to disease.
Sankaran began to look for commonalities across the DNA of people with sickle cell disease who had fetal hemoglobin, compared to those who didn’t. At the same time, a group led by Antonio Cao in Italy was already a step ahead, and had identified a few spots linked to the fetal protein using the tactic, known as a genome-wide association study, or GWAS.
In 2007, Cao submitted the study to the New England Journal of Medicine, but the editors rejected it, asking for a second cohort for confirmation. Cao’s lab teamed up with Orkin’s group, and with their sickle cell study, linked a gene called BCL11A to persistent production of fetal hemoglobin in adults.
“The journal still rejected it,” Orkin said. “I have absolutely no idea why.”
But Sankaran had already moved on to his next, and the most crucial discovery: that BCL11A was cranked up in adult blood cells, but not fetal ones. When he used a synthetic RNA molecule to block its production in blood stem cells, fetal hemoglobin shot up.
The key to treating sickle cell with fetal hemoglobin wasn’t a complex constellation of genes. It was the single switch they’d been seeking. Sankaran was working in the lab late that night and excitedly called Orkin to tell him the results. The implications were immediate. After decades of dead ends, the holy grail of hematology was finally found.
A selective strategy
The landmark results were published in Science in 2008 — next to the short mention of CRISPR. Sankaran left to finish medical school, and an incoming research fellow, Dan Bauer, would carry the torch and help turn the discovery into a therapeutic strategy.
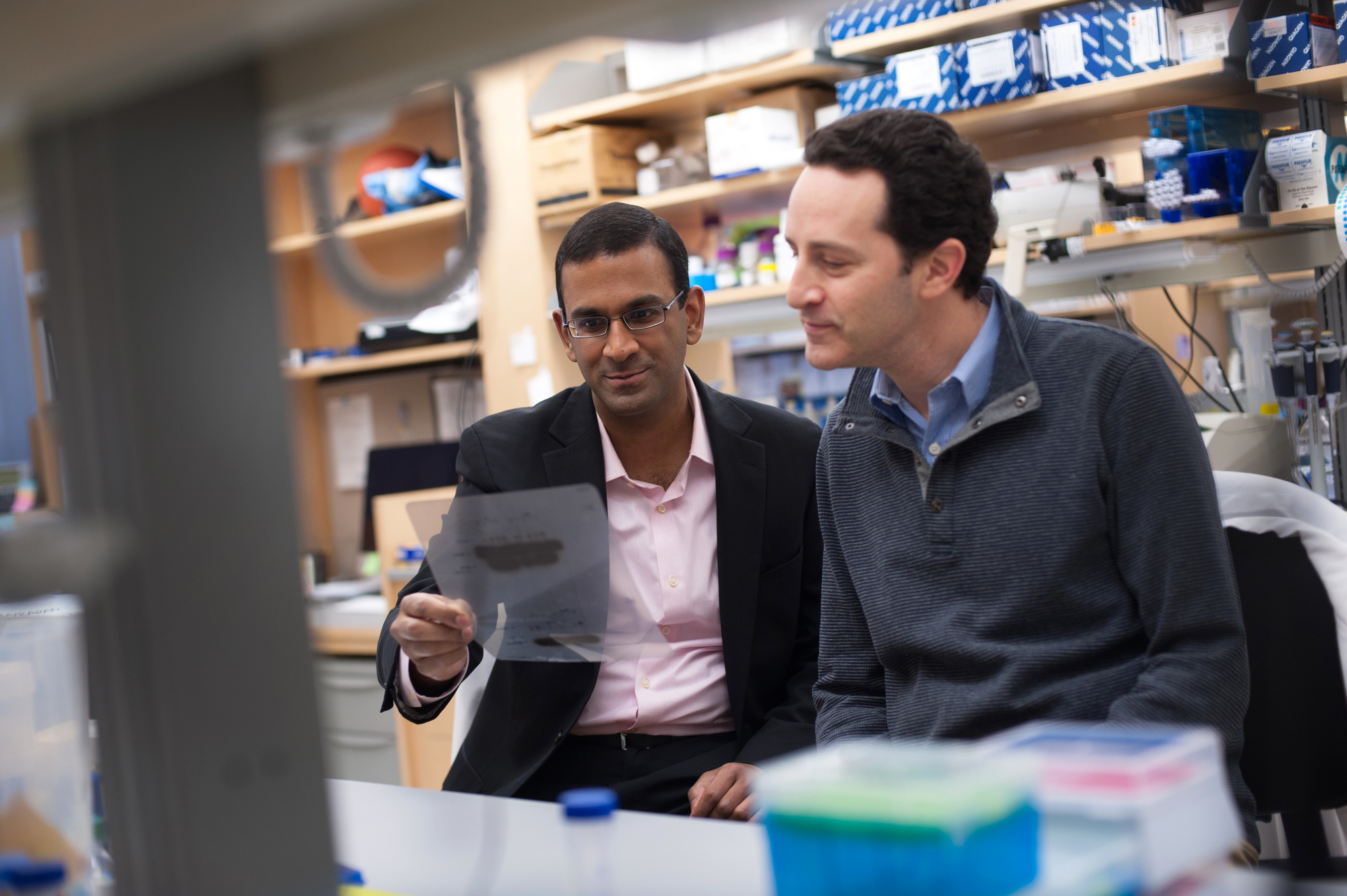
Vijay Sankaran (L) and Dan Bauer (Boston Children’s Hospital)
Click on the image to see the full-sized version
Doing so would prove tricky. Breaking BCL11A in all cells would cause problems — bone marrow cells, for example, require the protein to give rise to red blood cells and other things circulating throughout the body.
They needed a way to shut the switch off only in red blood cells. But the variants they needed to target were buried in part of the gene called an enhancer, long lines of code that regulate the production of the switch itself.
Using an older and more laborious gene-editing technology known as TALENs, Bauer removed the enhancer, which triggered an almost complete loss of BCL11A in red blood cells, while sparing other cells, and boosting fetal hemoglobin in the process.
“It was a much bigger effect than I expected,” Bauer said.
Fyodor Urnov, a gene editing scientist who was working on sickle cell disease at Sangamo Therapeutics at the time, called Bauer and Orkin’s work a “slightly mystical moment” for the field.
“That gave us a path towards interfering with BCL11A function without any side effects,” said Urnov, who is now a director at the Innovative Genomics Institute at UC Berkeley and a consultant to Vertex. “If you had told me 15 years ago that we would be gene editing an enhancer to treat sickle cell disease, I would have politely questioned your sanity.”
Early adopters
As Bauer’s enhancer paper was going to press in 2013, CRISPR was moving from what had been a curiosity in the 2008 pages of Science, into a revolutionary new technology. Bauer figured if they could break the enhancer with a key snip from the molecular scissors at just the right spot, it would be a simpler and safer approach than yanking the whole thing out.
Orkin, now 77, was one of CRISPR’s early adopters and partnered with one of the technology’s inventors, Feng Zhang at MIT, to figure out where precisely within the enhancer’s 13,000 letters of code to target.
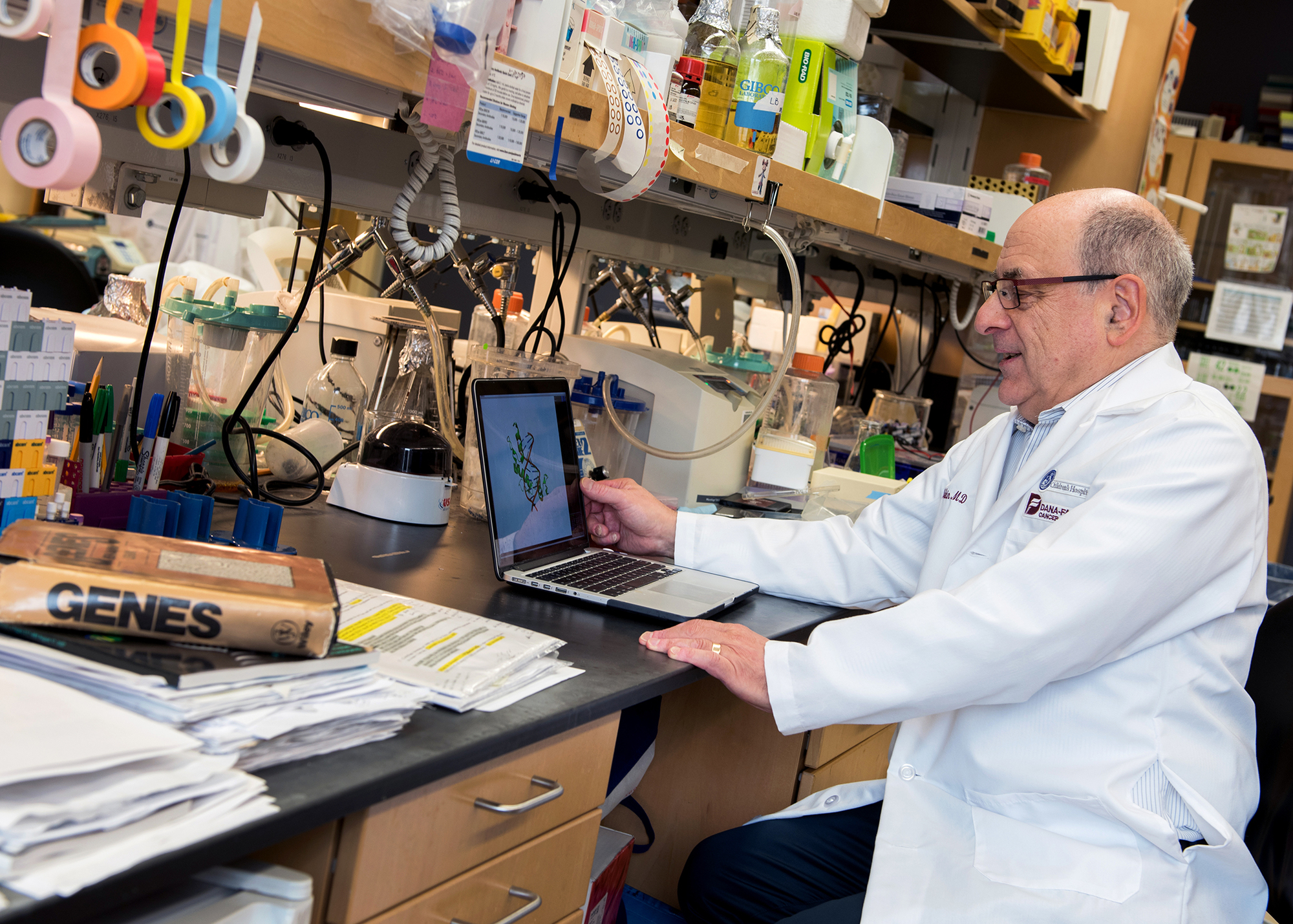
Stuart Orkin in 2017 (Boston Children’s Hospital)
Click on the image to see the full-sized version
The team published in Nature in 2015 and filed patents on several guide RNA molecules that are used to direct CRISPR to the BCL11A enhancer sites.
But Orkin said he wasn’t interested in pushing the experimental therapy into clinical trials on his own.
“If you really want to bring it to patients, it’s got to be done by pharma,” he said. “If we had started a trial here, we’d still be on our third patient or something.”
He instructed the hospital to make licenses to the technology non-exclusive, a decision that would result in smaller royalties for Boston Children’s, but would maximize potential access to the many companies eyeballing gene editing therapies for sickle cell at the time, including CRISPR, Editas, Intellia, Sanagmo and Vertex.
“I didn’t want to bet on one,” Orkin said.
In the meantime, Altshuler — the genetic professor whose lectures inspired Sankaran’s work — had ended up at Vertex. He’d shared an office with Zhang in his previous role at the Broad Institute of MIT and Harvard, was attuned to the potential of gene editing, and helped strike a partnership with CRISPR Therapeutics that fall.
CRISPR and Vertex looked for their own guide molecules to go after BCL11A, but ultimately ended up using one of Orkin’s.
“They couldn’t find a better one,” Orkin said. “What they’ve done is directly translate what we described.”
Less than four years later, 34-year-old patient Victoria Gray got an infusion of her own blood stem cells, carefully snipped in the BCL11A enhancer to boost fetal hemoglobin, in hopes that it would finally put an end to the excruciating pain crises from her sickled cells.
A qualified landmark
The rest is history. Nearly all sickle cell patients who have gotten Casgevy in clinical trials remain largely free of pain crises, and most beta thalassemia patients no longer require regular blood transfusions.
“There’s no question it’s a landmark. The question is if it’s more important as an example of editing, or as a good therapy for this disease?” Orkin said. “The results are great. But it’s going to be hard to impact the disease, because there are lots of people that will never get treated.”
These therapies require removing stem cells from a patient’s bone marrow, editing the cells in a lab, and using chemotherapy to wipe out a patient’s diseased cells to make room for the altered ones. It’s an arduous, time-consuming, and costly approach that means these therapies are unlikely to make a dent in the lower-income countries where sickle cell disease is more prevalent.
In the meantime, Orkin is working on a different approach: a pill that blocks BCL11A. “It does seem kind of backwards,” he admits.
It’s a challenge. BCL11A belongs to a class of proteins called transcription factors that are notoriously difficult to drug, never mind the headache of getting the compound to target red blood cells and not their stem cell precursors.
“If you tell me something’s undruggable I want to see if I can drug it. The challenge is part of it,” Orkin said. “It’s probably too hard. But I’m having a reasonably good time trying to figure it out.”

