Unraveling the Complexities of Cancer through More Human-relevant Models
The current paradigm in preclinical testing for developing cancer therapeutics is failing. Oncology drug candidates are currently the least likely type of therapeutic to succeed in clinical trials, with only 5.1% of Phase I candidates going on to receive FDA approval. A major reason for this high attrition rate is the preclinical in vitro and in vivo models that cancer researchers rely on. While conventional in vitro cancer models can mimic some parts of human physiology, they often lack critical factors such as cellular heterogeneity and a dynamic, native microenvironment. As such, scientists continue to rely on animal models; however, animals often lack human relevance, allowing ineffective and toxic drugs to enter clinical trials. To create more successful drug development programs, scientists need models that provide a more comprehensive and human-relevant view of cancer progression.
The Importance of Understanding the Tumor Microenvironment
There is growing evidence that the tumor microenvironment (TME) is a crucial modulator of cancer growth, migration, angiogenesis, immunosurveillance evasion, and therapy resistance. The TME is composed of extracellular matrix (ECM) and various non-malignant cell types, including endothelial, mesenchymal (e.g., fibroblasts and adipocytes) and immune cells (e.g., lymphocytes, monocytes, and neutrophils), all of which interact with tumor cells via secretion of molecules such as growth factors, cytokines, extracellular vesicles, and miRNAs (Figure 1). The ECM in particular has a significant effect on the behavior of cells, as it provides structural and biochemical support. During cancer progression, the stiffness of the ECM influences cells’ proliferation, survival, migration, and differentiation. Thus, understanding the TME is key to understanding overall cancer progression and developing effective therapies.
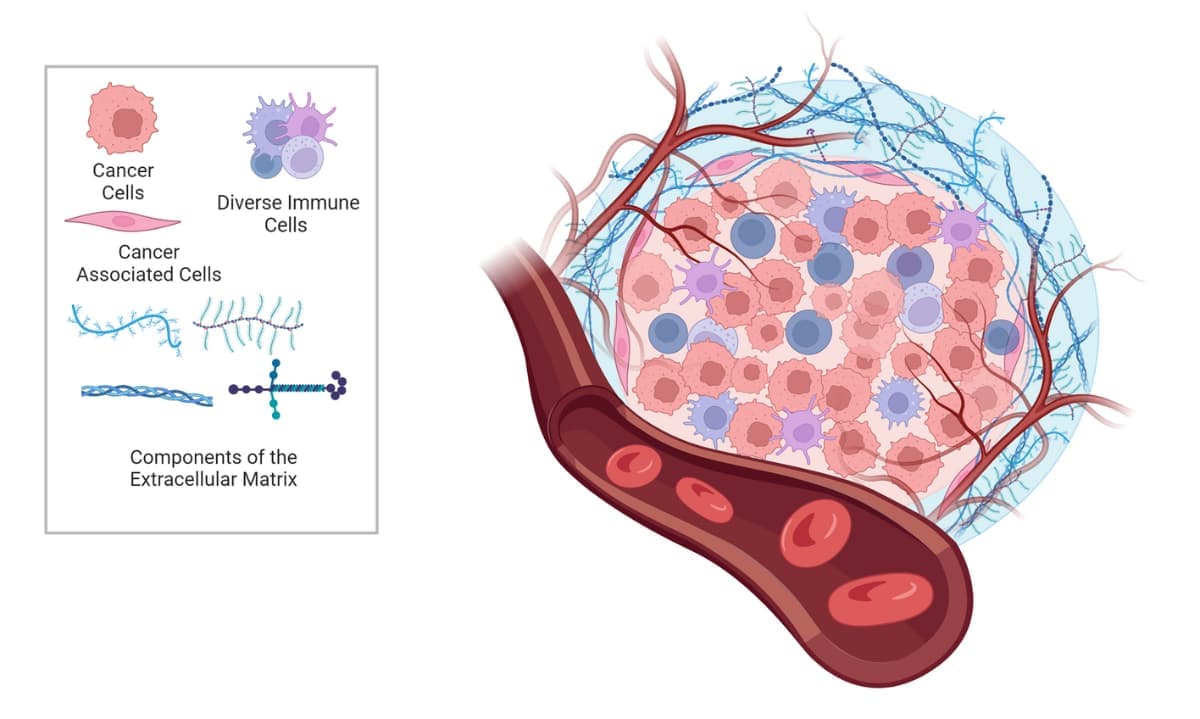
Figure 1. The tumor microenvironment.
The Need for Better Models
Animal models have traditionally been considered the gold standard in disease research and drug development, despite the high failure rate of drugs transitioning to the clinic. To improve the translatability of animal models for cancer research, human tumors are often grafted into animals. The xenograft mouse model, for example, is created when the tissue of interest is implanted into an immunocompromised mouse. The orthotopic mouse model differs in that the human tumor is grafted onto the corresponding organ of interest in the mouse. While these models are more difficult to create, they are more physiologically relevant because the tumors are growing in an organ-specific TME. Despite using human tumors, however, xenograft and orthotopic mouse models still have limited human relevance, as the immune components and TME are animal-based and do not recapitulate how a tumor progresses in the human body.
While researchers can create cancer models entirely from human cell sources, they continue to face challenges in fully recapitulating human biological function in a dish. 2D in vitro models are simple and cost-effective models to incorporate into drug screening assays, but their simplicity results in low physiological relevance. Spheroids offer improved 3D cytoarchitecture, but often have issues in uniformity and reproducibility, lack standardized assessments of growth and drug efficacy, and are not high throughput for drug screening.
Organoids are a powerful alternative to 2D culture that preserves many of the structural and functional traits of their in vivo counterparts, such as a hypoxic microenvironment, cell heterogeneity, ECM interactions, and more in vivo-relevant gene expression patterns. Tumor organoids can either be derived from patient tissue or engineered using induced pluripotent stem cells (iPSCs) and special culture conditions. Those generated from iPSCs and adult stem cells encompass important tissue features such as architecture, differentiated cell types, and tissue function. Taken together, organoids are comparable to certain in vivo models such as traditional genetically engineered mouse models, cell lines, and patient-derived xenografts (PDX). However, organoids lack an organ-specific environment that includes immune components, blood vessels, and different stromal cells. Finally, growth stimulators and inhibitors used in the growth of organoids can affect drug sensitivity, gene expression, and cell signaling pathways.
A New Way Forward: Improving 3D Cancer Modeling with Organ-Chips
Organ-on-a-Chip technology has been developed to address many of the challenges discussed above. These microfluidic devices emulate in vivo physiology and tissue microenvironments in an organ-specific context by enabling key cellular populations to be added to parallel microfluidic channels that are separated by a porous membrane (Figure 2). Additional features such as biomechanical forces, ECM, and immune cells provide a more human-relevant environment for cells to behave as they would in vivo.
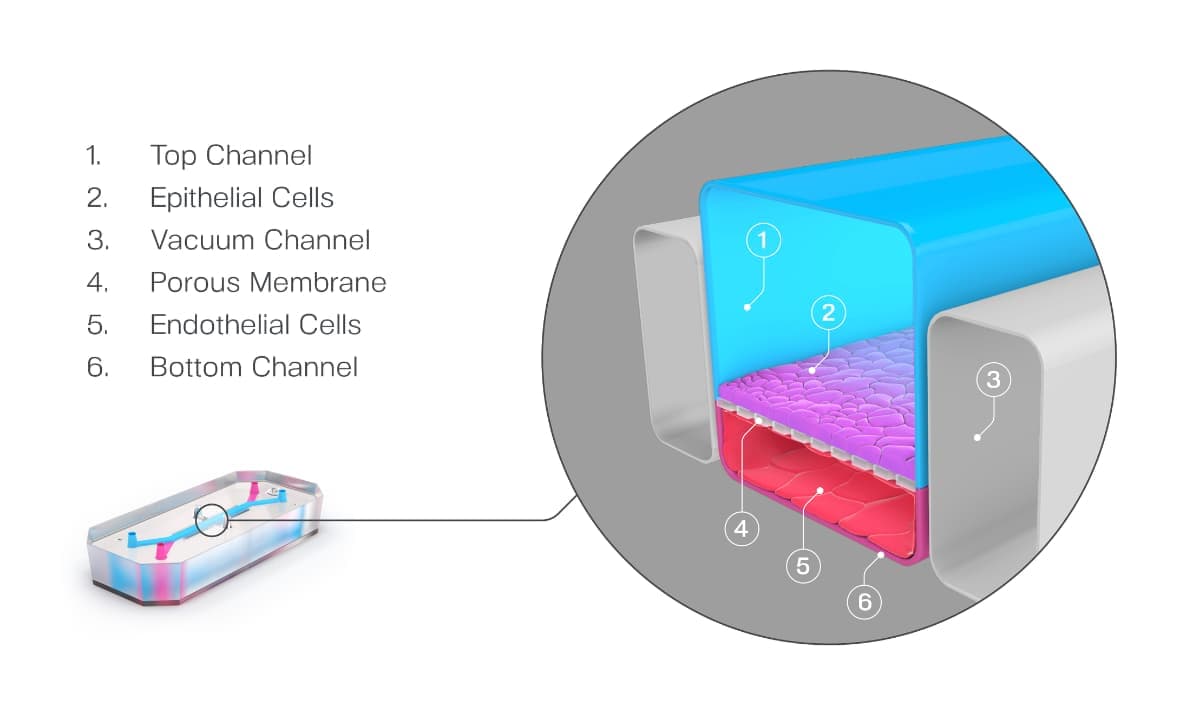
Figure 2. Schematic of an Organ-Chip.
Recent advances in the design of Organ-Chip consumables have improved the ability of Organ-Chips to address the needs of modeling the tumor microenvironment. The Emulate Chip-A1 Accessible Chip allows researchers to model complex 3D tissues by incorporating gels up to 3 mm thick within the chip’s accessible culture chamber, integrating stroma into the epithelial layer, creating stratified epithelia, and applying organ-specific biomechanical forces (Figure 3). Further complexity and human relevance can be achieved with Chip-A1 by incorporating circulating immune cells such as PBMCs or CAR T cells.
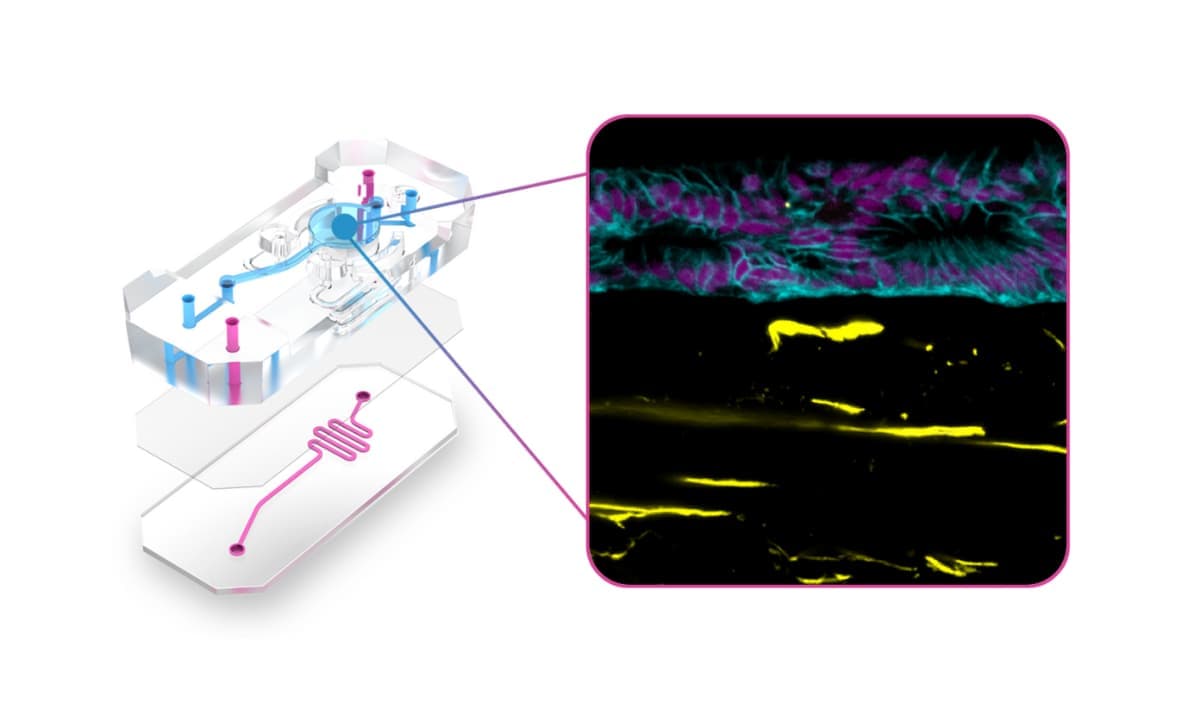
Figure 3: Exploded view of Chip-A1 Accessible Chip demonstrating how it has been used to model epithelial-stromal interactions in Barrett’s Esophagus.
Chip-A1 has already enabled researchers to better understand how the surrounding microenvironment influences cancer progression. In their online article titled ‘Epithelial-Stromal Interactions in Barrett’s Esophagus Modeled in Human Organ Chips,” researchers from the Wyss Institute for Biologically Inspired Engineering at Harvard University used a prototype of Chip-A1 and found that it offered a new approach for studying epithelial-stromal interactions and the broader underlying mechanisms associated with esophageal cancer progression. The team also reported that this model could potentially serve as a tool for personalized drug-response assessments between different patients or genetic subpopulations.
Conclusion
Understanding how the tumor microenvironment contributes to cancer progression remains critical for developing more effective therapeutics. Organ-on-a-Chip technology is evolving to meet the in vitro modeling needs for research areas such as cancer, which require enhanced 3D models and more physiologically relevant compound dosing options. Preclinical models that enable researchers to model biology that is more complex, like the TME, will improve clinical translation, and Chip-A1 will give researchers this capability.